Strontium »
PDB 1cs7-2grb »
2awe »
Strontium in PDB 2awe: Base-Tetrad Swapping Results in Dimerization of Rna Quadruplexes: Implications For Formation of I-Motif Rna Octaplex
Protein crystallography data
The structure of Base-Tetrad Swapping Results in Dimerization of Rna Quadruplexes: Implications For Formation of I-Motif Rna Octaplex, PDB code: 2awe
was solved by
B.Pan,
K.Shi,
M.Sundaralingam,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 19.36 / 2.10 |
| Space group | I 41 2 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 82.667, 82.667, 74.154, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 17.2 / 22.8 |
Other elements in 2awe:
The structure of Base-Tetrad Swapping Results in Dimerization of Rna Quadruplexes: Implications For Formation of I-Motif Rna Octaplex also contains other interesting chemical elements:
| Bromine | (Br) | 8 atoms |
| Potassium | (K) | 11 atoms |
Strontium Binding Sites:
The binding sites of Strontium atom in the Base-Tetrad Swapping Results in Dimerization of Rna Quadruplexes: Implications For Formation of I-Motif Rna Octaplex
(pdb code 2awe). This binding sites where shown within
5.0 Angstroms radius around Strontium atom.
In total only one binding site of Strontium was determined in the Base-Tetrad Swapping Results in Dimerization of Rna Quadruplexes: Implications For Formation of I-Motif Rna Octaplex, PDB code: 2awe:
In total only one binding site of Strontium was determined in the Base-Tetrad Swapping Results in Dimerization of Rna Quadruplexes: Implications For Formation of I-Motif Rna Octaplex, PDB code: 2awe:
Strontium binding site 1 out of 1 in 2awe
Go back to
Strontium binding site 1 out
of 1 in the Base-Tetrad Swapping Results in Dimerization of Rna Quadruplexes: Implications For Formation of I-Motif Rna Octaplex
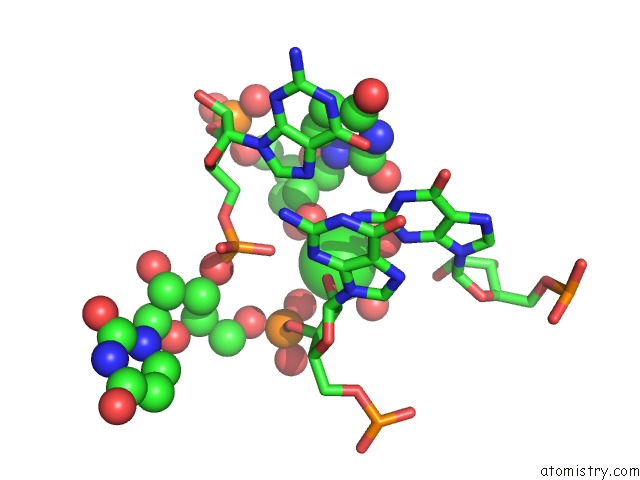
Mono view
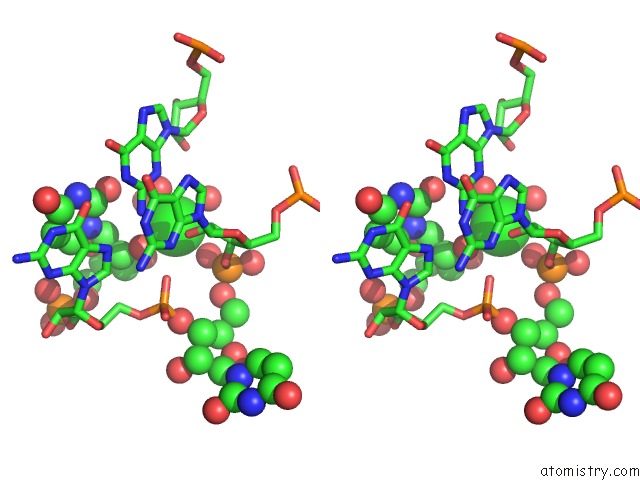
Stereo pair view

Mono view

Stereo pair view
A full contact list of Strontium with other atoms in the Sr binding
site number 1 of Base-Tetrad Swapping Results in Dimerization of Rna Quadruplexes: Implications For Formation of I-Motif Rna Octaplex within 5.0Å range:
|
Reference:
B.Pan,
K.Shi,
M.Sundaralingam.
Base-Tetrad Swapping Results in Dimerization of Rna Quadruplexes: Implications For Formation of the I-Motif Rna Octaplex. Proc.Natl.Acad.Sci.Usa V. 103 3130 2006.
ISSN: ISSN 0027-8424
PubMed: 16492787
DOI: 10.1073/PNAS.0507730103
Page generated: Thu Oct 10 20:55:59 2024
ISSN: ISSN 0027-8424
PubMed: 16492787
DOI: 10.1073/PNAS.0507730103
Last articles
Ca in 5S5ICa in 5S5H
Ca in 5S5G
Ca in 5S5F
Ca in 5S5D
Ca in 5S5E
Ca in 5S5C
Ca in 5S5B
Ca in 5S5A
Ca in 5S59