Strontium »
PDB 3ilg-4rne »
4o41 »
Strontium in PDB 4o41: Amide Linked Rna
Protein crystallography data
The structure of Amide Linked Rna, PDB code: 4o41
was solved by
P.S.Pallan,
M.Egli,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 28.00 / 1.20 |
| Space group | P 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 22.360, 34.330, 43.300, 110.03, 94.24, 95.83 |
| R / Rfree (%) | 14.7 / 21.2 |
Strontium Binding Sites:
The binding sites of Strontium atom in the Amide Linked Rna
(pdb code 4o41). This binding sites where shown within
5.0 Angstroms radius around Strontium atom.
In total 2 binding sites of Strontium where determined in the Amide Linked Rna, PDB code: 4o41:
Jump to Strontium binding site number: 1; 2;
In total 2 binding sites of Strontium where determined in the Amide Linked Rna, PDB code: 4o41:
Jump to Strontium binding site number: 1; 2;
Strontium binding site 1 out of 2 in 4o41
Go back to
Strontium binding site 1 out
of 2 in the Amide Linked Rna

Mono view
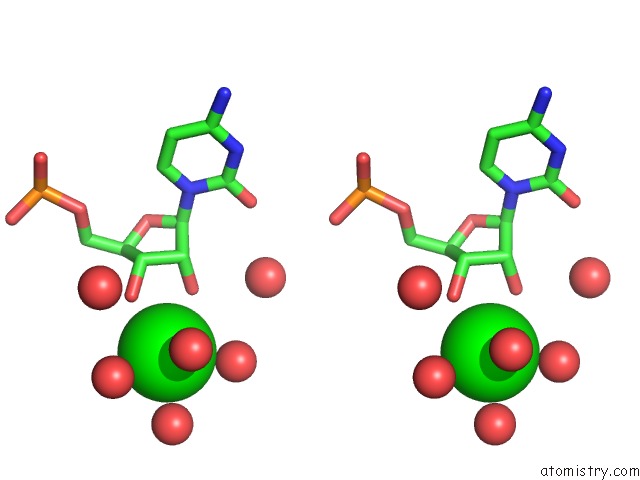
Stereo pair view

Mono view

Stereo pair view
A full contact list of Strontium with other atoms in the Sr binding
site number 1 of Amide Linked Rna within 5.0Å range:
|
Strontium binding site 2 out of 2 in 4o41
Go back to
Strontium binding site 2 out
of 2 in the Amide Linked Rna

Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Strontium with other atoms in the Sr binding
site number 2 of Amide Linked Rna within 5.0Å range:
|
Reference:
D.Mutisya,
C.Selvam,
B.D.Lunstad,
P.S.Pallan,
A.Haas,
D.Leake,
M.Egli,
E.Rozners.
Amides Are Excellent Mimics of Phosphate Internucleoside Linkages and Are Well Tolerated in Short Interfering Rnas. Nucleic Acids Res. V. 42 6542 2014.
ISSN: ISSN 0305-1048
PubMed: 24813446
DOI: 10.1093/NAR/GKU235
Page generated: Fri Oct 11 05:44:24 2024
ISSN: ISSN 0305-1048
PubMed: 24813446
DOI: 10.1093/NAR/GKU235
Last articles
Ca in 2VK6Ca in 2VJ3
Ca in 2VJJ
Ca in 2VHD
Ca in 2VDR
Ca in 2VH0
Ca in 2VFQ
Ca in 2VFO
Ca in 2VFP
Ca in 2VFN